The LwCas13a/Cas13a expression plasmid is now available here!>>
In a study ("Nucleic Acid Detection with CRISPR-Cas13a/C2c2") published in Science, Broad institute investigators describe how this RNA-targeting CRISPR enzyme was harnessed as a sensitive detector that was able to indicate the presence of as little as a single molecule of a target RNA or DNA molecule. The new tool, dubbed SHERLOCK (Specific High-sensitivity Enzymatic Reporter unLOCKing), could one day be used to respond to viral and bacterial outbreaks, monitor antibiotic resistance, and detect cancer, according to the researchers.
Here we presented the sequence of Cas13a(previously known as C2c2), also with the expression and purification of the Leptotrichia wadei (LwCas13a).
The sequence of the LwCas13a
The backbond of the plasmid used for expression and pufication of LwCas13a in the paper is pET vector which is deriven by T7 promoter in E.coli. The plasmid map of the plasmid(pET His6-TwinStrep-SUMO-LwCas13a) is as below(please see the full length sequence of the plasmid at the end of the article):
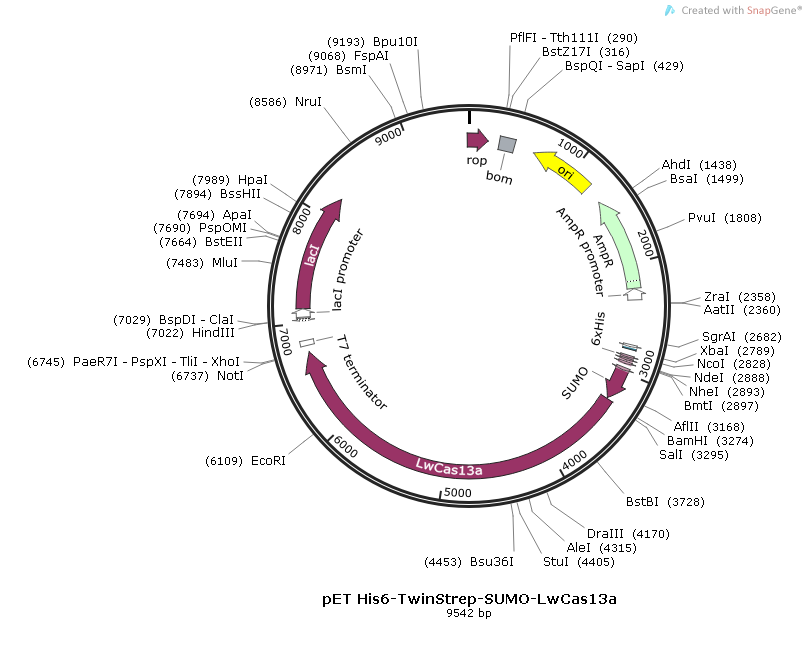
LwCas13a protein purification
Cas13a bacterial expression vectors were transformed into Rosetta™ 2(DE3)pLysS Singles Competent Cells (Millipore). A 16 mL starter culture was grown overnight in Terrific Broth 4 growth media (12 g/L tryptone, 24 g/L yeast extract, 9.4 g/L K2HPO, 2.2 g/L KH2PO4, Sigma) (TB), which was used to inoculate 4 L of TB for growth at 37°C and 300 RPM until an OD600 of 0.6.
At this time, protein expression was induced by supplementation with IPTG (Sigma) to a final concentration of 500 uM, and cells were cooled to 18°C for 16 h for protein expression. Cells were then centrifuged at 5200 g for 15 min at 4°C. Cell pellet was harvested and stored at - 80°C for later purification.
All subsequent steps of the protein purification were performed at 4°C. Cell pellet was crushed and resuspended in lysis buffer (20 mM Tris-HCl, 500 mM NaCl, 1 mM DTT, pH 8.0) supplemented with protease inhibitors (Complete Ultra EDTA-free tablets), lysozyme, and benzonase followed by sonication (Sonifier 450, Branson, Danbury, CT) with the following conditions: amplitude of 100 for 1 second on and 2 seconds off with a total sonication time of 10 min. Lysate was cleared by centrifugation for 1 hr at 4°C at 10,000 g and the supernatant was filtered through a Stericup 0.22 μm filter (EMD Millipore).
Filtered supernatant was applied to StrepTactin Sepharose (GE) and incubated with rotation for 1 hr followed by washing of the protein-bound StrepTactin resin three times in lysis buffer. The resin was resuspended in SUMO digest buffer (30 mM Tris-HCl, 500 mM NaCl 1 mM DTT, 0.15% Igepal (NP-40), pH 8.0) along with 250 Units of SUMO protease (ThermoFisher) and incubated overnight at 4°C with rotation. Digestion was confirmed by SDS-PAGE and Coomassie Blue staining and the protein eluate was isolated by spinning the resin down.
For further cation exchange and gel filtration purification, protein was loaded onto a 5 mL HiTrap SP HP cation exchange column (GE Healthcare Life Sciences) via FPLC (AKTA PURE, GE Healthcare Life Sciences) and eluted over a salt gradient from 130 mM to 2M NaCl in elution buffer (20 mM Tris-HCl, 1 mM DTT, 5% glycerol, pH 8.0).
The resulting fractions were tested for presence of LwCas13a by SDS-PAGE, and fractions containing the protein were pooled and concentrated via a Centrifugal Filter Unit to 1 mL in S200 buffer (10 mM HEPES, 1 M NaCl, 5 mM MgCl2, 2 mM DTT, pH 7.0). The concentrated protein was loaded onto a gel filtration column (Superdex® 200 Increase 10/300 GL, GE Healthcare Life Sciences) via FPLC.
The resulting fractions from gel filtration were analyzed by SDS-PAGE and fractions containing LwCas13a were pooled and buffer exchanged into Storage Buffer (600 mM NaCl, 50 mM Tris- HCl pH 7.5, 5% glycerol, 2mM DTT) and frozen at -80°C for storage.
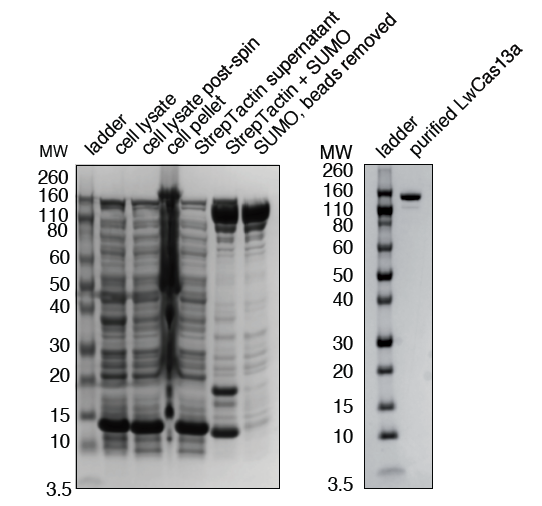
Coomassie blue stained acrylamide gel of LwCas13a stepwise purification.
The full length of plasmid pET His6-TwinStrep-SUMO-LwCas13a
All data were cited from: Gootenberg JS, Abudayyeh OO, Lee JW, et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science. 2017.
Souce: NovoPro 2017-04-16
